Model info
| Transcription factor | Tp53 | ||||||||
| Model | P53_MOUSE.H10MO.C | ||||||||
| Model type | Mononucleotide PWM | ||||||||
| LOGO | 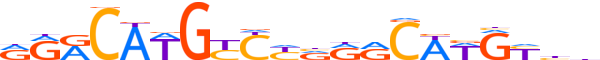 | ||||||||
| LOGO (reverse complement) |  | ||||||||
| Origin | HOCOMOCO v9 | ||||||||
| Model length | 20 | ||||||||
| Quality | C | ||||||||
| Consensus | RRRCAWGYCYvRRCWWGYhb | ||||||||
| wAUC | |||||||||
| Best AUC | |||||||||
| Benchmark datasets | |||||||||
| Aligned words | 1754 | ||||||||
| TF family | p53-related factors{6.3.1} | ||||||||
| TF subfamily | p53{6.3.1.0.1} | ||||||||
| MGI | 98834 | ||||||||
| EntrezGene | 22059 | ||||||||
| UniProt ID | P53_MOUSE | ||||||||
| UniProt AC | P02340 | ||||||||
| Comment | |||||||||
| Downloads | pcm
pwm alignment threshold to P-value grid | ||||||||
| Standard thresholds |
|
PCM
| A | C | G | T | |
|---|---|---|---|---|
| 01 | 508.352 | 51.592 | 921.141 | 127.533 |
| 02 | 280.937 | 37.611 | 1187.12 | 102.95 |
| 03 | 859.005 | 23.21 | 710.184 | 16.218 |
| 04 | 9.251 | 1563.55 | 34.209 | 1.608 |
| 05 | 1393.858 | 18.106 | 46.84 | 149.814 |
| 06 | 325.467 | 9.018 | 79.184 | 1194.949 |
| 07 | 5.336 | 2.144 | 1600.601 | 0.536 |
| 08 | 50.334 | 784.539 | 32.112 | 741.633 |
| 09 | 92.65 | 1306.41 | 65.48 | 144.078 |
| 10 | 154.354 | 857.395 | 42.644 | 554.225 |
| 11 | 269.025 | 224.614 | 953.912 | 161.067 |
| 12 | 353.988 | 70.002 | 1038.591 | 146.037 |
| 13 | 600.884 | 48.657 | 885.37 | 73.707 |
| 14 | 59.47 | 1444.19 | 91.675 | 13.283 |
| 15 | 1215.896 | 91.464 | 72.239 | 229.018 |
| 16 | 255.724 | 92.582 | 131.987 | 1128.324 |
| 17 | 128.071 | 91.814 | 1335.625 | 53.108 |
| 18 | 184.626 | 304.704 | 88.435 | 1030.853 |
| 19 | 239.269 | 646.702 | 169.692 | 552.954 |
| 20 | 234.184 | 526.536 | 276.233 | 571.665 |
PWM
| A | C | G | T | |
|---|---|---|---|---|
| 01 | 0.233 | -2.023 | 0.826 | -1.139 |
| 02 | -0.357 | -2.326 | 1.079 | -1.349 |
| 03 | 0.757 | -2.78 | 0.567 | -3.107 |
| 04 | -3.595 | 1.354 | -2.416 | -4.762 |
| 05 | 1.24 | -3.008 | -2.116 | -0.98 |
| 06 | -0.211 | -3.616 | -1.607 | 1.086 |
| 07 | -4.03 | -4.618 | 1.378 | -5.134 |
| 08 | -2.047 | 0.666 | -2.476 | 0.61 |
| 09 | -1.453 | 1.175 | -1.792 | -1.018 |
| 10 | -0.95 | 0.755 | -2.206 | 0.319 |
| 11 | -0.4 | -0.579 | 0.861 | -0.908 |
| 12 | -0.127 | -1.727 | 0.946 | -1.005 |
| 13 | 0.4 | -2.079 | 0.787 | -1.677 |
| 14 | -1.885 | 1.275 | -1.463 | -3.285 |
| 15 | 1.103 | -1.465 | -1.696 | -0.56 |
| 16 | -0.45 | -1.454 | -1.105 | 1.029 |
| 17 | -1.135 | -1.462 | 1.197 | -1.995 |
| 18 | -0.773 | -0.276 | -1.498 | 0.939 |
| 19 | -0.516 | 0.473 | -0.857 | 0.317 |
| 20 | -0.537 | 0.268 | -0.374 | 0.35 |